0 引言
1 材料与方法
1.1 样品采集
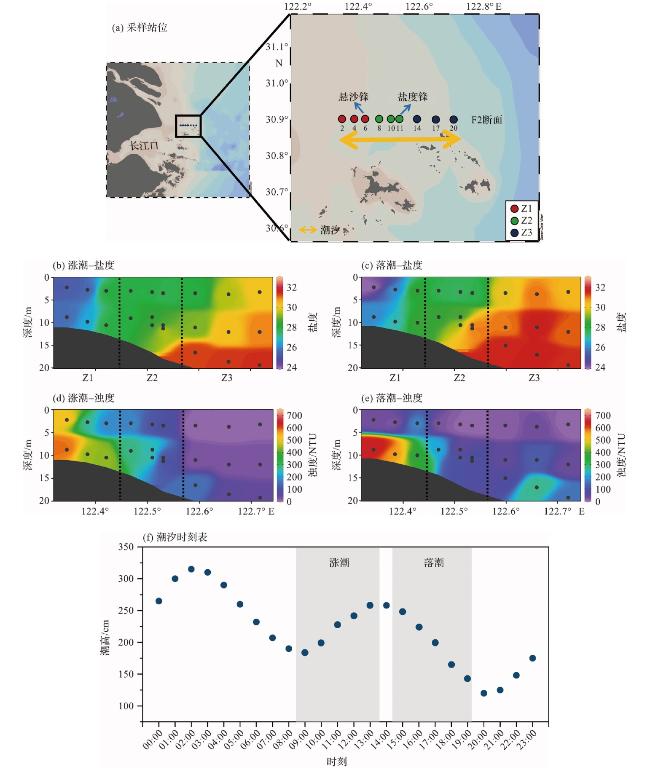
图1 长江口及其邻近海域采样站位分布与悬沙锋和盐度锋结构(站点F2-6和站点F2-11分别为悬沙锋和盐度锋。根据浊度和盐度特征,将研究区域分为Z1、Z2和Z3三个区。图1f中的阴影表示采样时间段。) Fig.1 Distribution of sampling stations and the structures of the turbidity front and salinity front in the Changjiang River Estuary and its adjacent waters (Sample sites F2-6 and F2-11 indicate the locations of turbidity front and salinity front, respectively. Based on turbidity and salinity characteristics,the study area was divided into Z1, Z2 and Z3 zones. The shaded areas in Figure 1f represent the sampling periods.) |
1.2 样品测定
1.2.1 DOC质量浓度的测定
1.2.2 环境参数的测定
1.3 代谢物样品提取
1.4 液相质谱仪分析方法
1.5 代谢组学数据分析
1.5.1 原始数据预处理
1.5.2 统计分析
2 结果
2.1 盐度和浊度的空间分布特征
表1 2023年长江口锋面海域浊度、盐度、温度的变化范围Tab.1 Variation ranges of turbidity, salinity, and temperature in the frontal sea area of the Changjiang River Estuary in 2023 |
| 参数 | 涨潮 | 落潮 | ||||
|---|---|---|---|---|---|---|
| 表层 | 中层 | 底层 | 表层 | 中层 | 底层 | |
| 浊度/NTU | 9.5~518.1 | 24.3~247.0 | 43.9~588.6 | 11.1~178.6 | 38.5~117.7 | 75.3~648.9 |
| 盐度 | 25.5~30.1 | 28.4~30.9 | 25.8~31.5 | 24.1~31.0 | 28.6~31.7 | 26.1~31.7 |
| 温度/℃ | 11.1~12.1 | 11.0~11.6 | 11.0~11.4 | 11.4~12.1 | 11.1~11.6 | 11.1~11.5 |
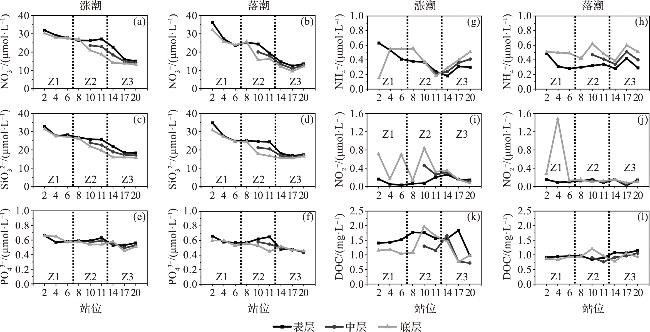
2.2 营养盐的空间分布特征
2.3 DOC的空间分布特征
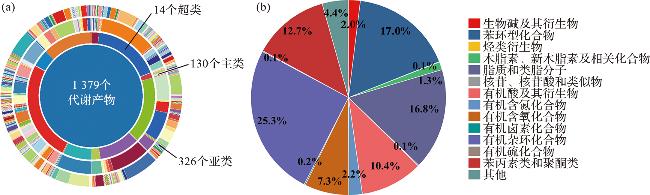
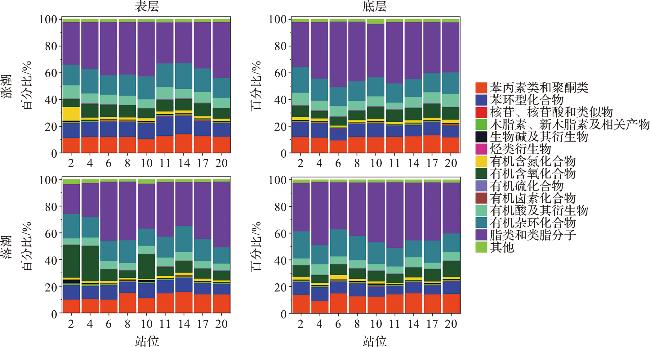
2.4 海水小分子代谢物的组成
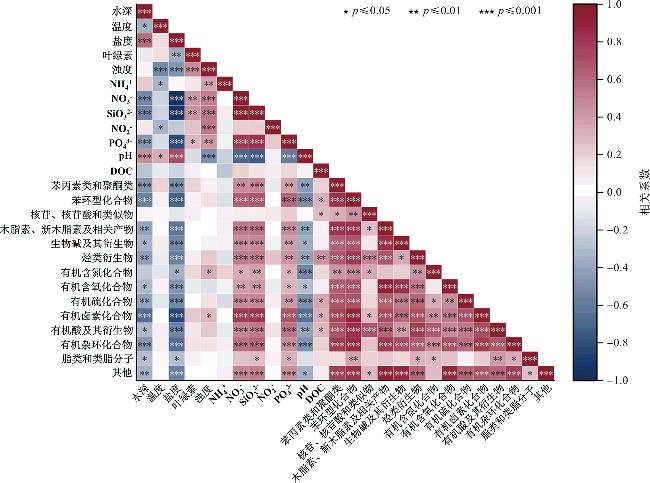
2.5 环境参数对代谢物在超类水平上的分布影响
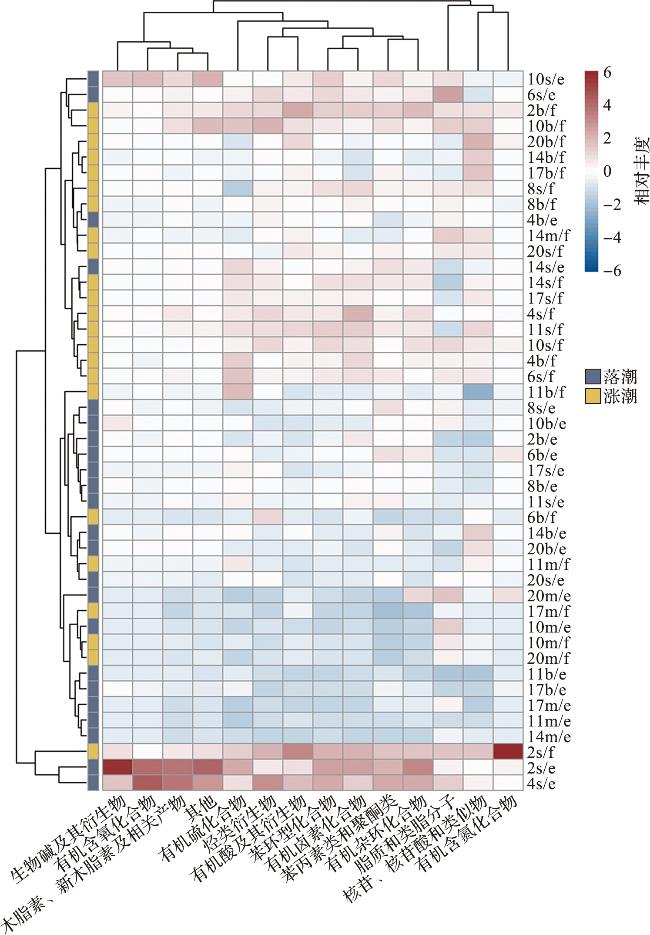
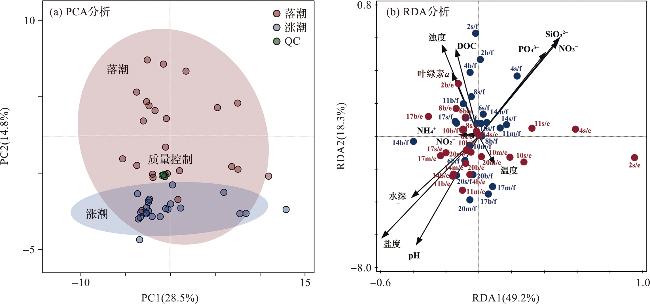
图6 海水溶解性代谢物组成分布的统计特征[图6b中的环境因子以箭头表示;海水被分为两组:涨潮(蓝点)和落潮(红点)。s:表层;m:中层;b:底层;f:涨潮;e:落潮。] Fig.6 Statistical characteristics of dissolved metabolite compositions in seawater [Environmental factors are represented by arrows, and seawater samples are divided into two groups: flood tide (blue dots) and ebb tide (red dots). s: surface; m: middle; b: bottom; f: flood; e: ebb.] |
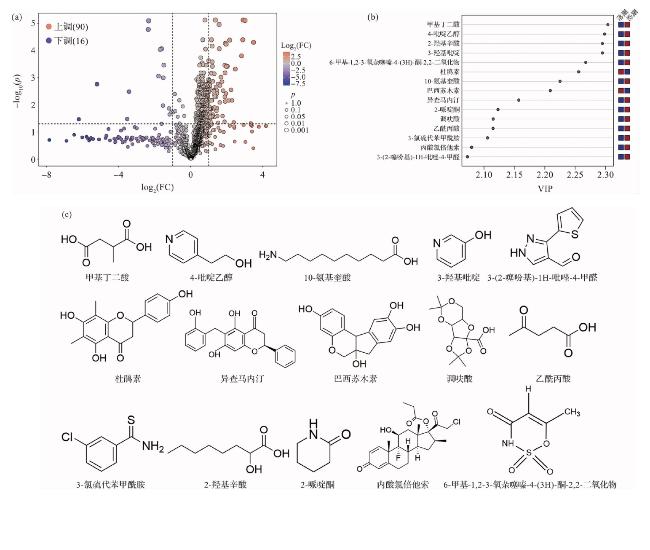
2.6 潮汐运动引起的代谢物差异化
图7 潮汐控制下的差异代谢物分析(a:涨、落潮时代谢物的变异倍数及其显著性p值;b:关键差异代谢物的投影变量重要性值(VIP)排序,红色方块表示含量相对高,蓝色方块表示含量相对低;c:VIP排名前15的代谢物分子结构。) Fig.7 Differential metabolites analysis under tidal influence (a: Fold change and significance p-values of metabolites during ebb and flood tides; b: Variable importance of the projection (VIP) scores of key differential metabolites,with red and blue squares referring the relatively high and low contents, respectively; c: Molecular structures of the top 15 metabolites ranked by VIP.) |